DELs in Cells – Snap – Seamless small molecule drug discovery – Get an early lead
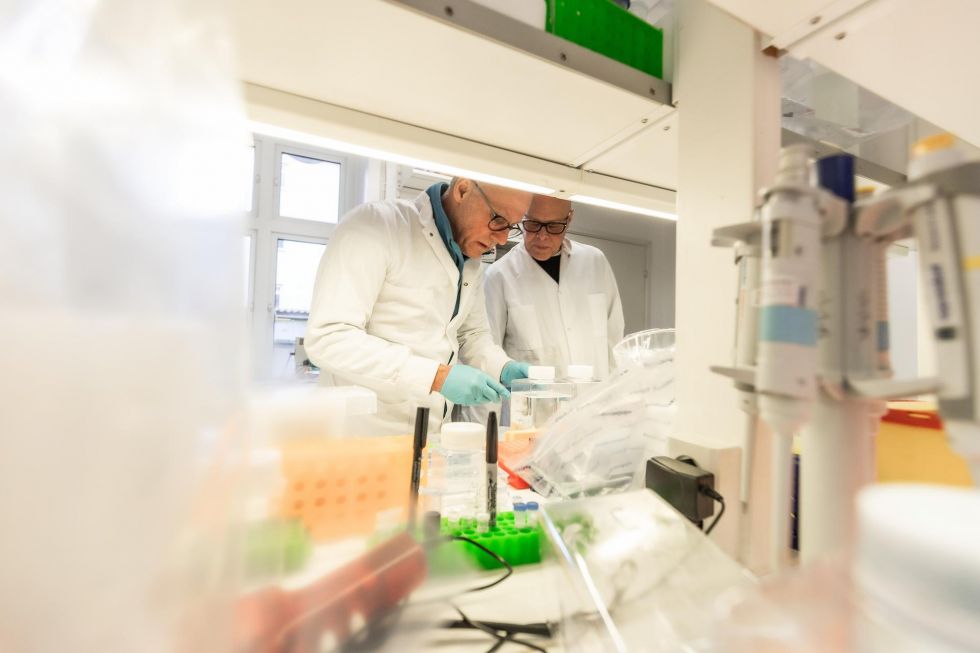
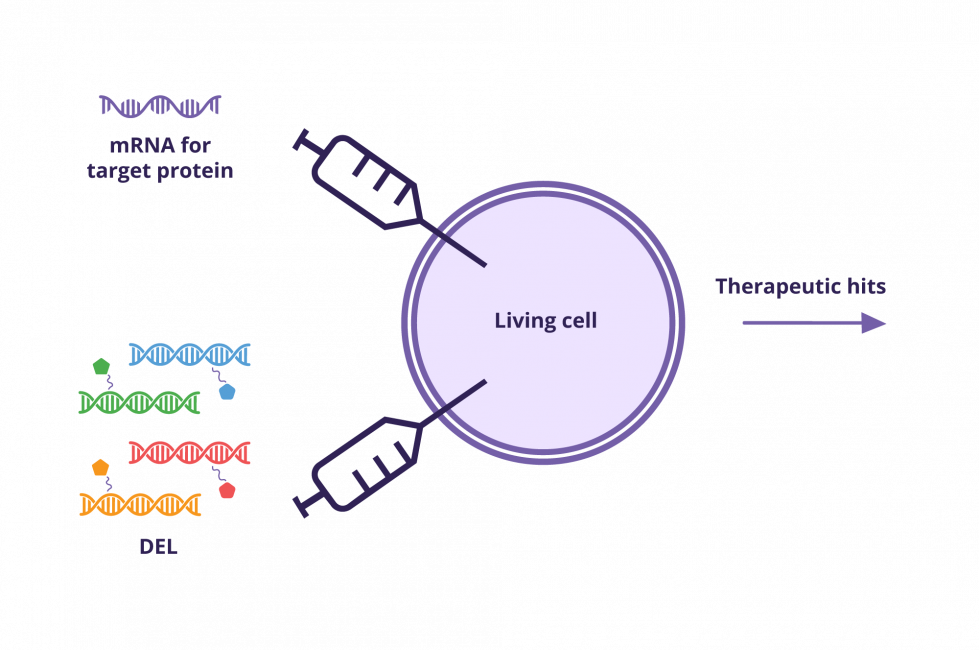
DELs in Cells – Snap. Our easy, fast, and affordable DNA-encoded chemical library (DEL) in living cells screening service.
Service in brief
-
1. You provide the amino acid sequence of the target protein(s)
-
2. We conduct an expression studyand provide a report
-
3. You say ”go”
-
4. We conduct the screening, perform the analysis and provide a report, and chemical structure information for 25 top hits
Timelines
Expression study: 4 weeks
Screening: 4 weeks
Keys
-
Screen is conducted in a living cell
-
Screen is conducted using 2 screening slots (2 constructs or 2 DELs) in parallel
-
Chemical structure information for 25 top hits
-
More physiologically relevant conditions
-
Lower attrition rate (assumed)
-
Broader target space – no need for purified target protein
-
High success rate (>70%)
-
Shorter TAT
-
Identifies hits and hit families
-
Low false positive rates (100% match between DNA code and compound)
-
Applicable across disease areas and target classes (no structural information required)
-
Straightforward and transparent data analysis
Business Model
- Simple fee-based plan with no reach-through and royalty payments
- Hit exclusivity and transparent cost structure – Client owns all hits and foreground IP without any downstream financial obligations
- The compensation plan comprises a total of 3 payments as a transparent cost structure
- Pre-Project Expression Study Fee – scales with work
- Screening Fee – scales with screening slots
- Chemical Series Nomination Fee
- The project flow is rapid and straightforward:
- Client provides the amino acid sequence for the target protein
- Vipergen performs the Expression Study (go-no-go)
- Vipergen performs one round of screenings with varying constructs and different libraries.
- Vipergen delivers a hit list and report, including metrics from screens and chemical structure information
- Hits are reserved for client for 1 year
- Client may resynthesize and test hits in relevant assays and use data for 1 year
- Vipergen can assist in off-DNA resynthesis as a fee-for-service
- Client may nominate Chemical Series for 1 year
- Vipergen assigns its rights to Nominated Chemical Series, and these become exclusive for you as a client
- Client own all foreground IP, data and hits of nominated series without royalty payment or other downstream financial obligations
DELs
Vipergen’s high fidelity DELs are synthesized using robust chemistry and with a purification step after each chemical transformation, thus providing 100% correspondence between code and compound (no truncates).
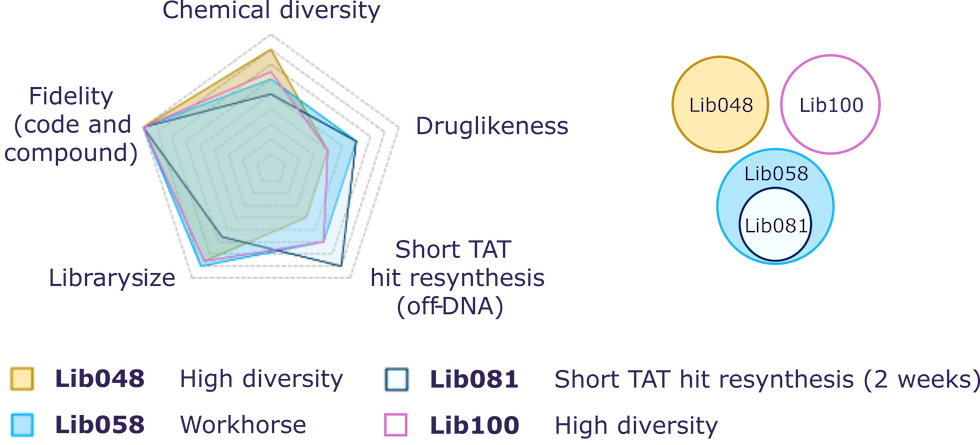
Key drivers for library design
- Chemical diversity
- Physicochemical properties
- Fidelity
- Hit resynthesis (off DNA)
| Lib048 | Lib058 | Lib081 | Lib100 | |
|---|---|---|---|---|
| Size & Diversity | ||||
| Size (million compounds) | 551 | 603 | 381 | 511 |
| Building blocks | 1616 | 1339 | 1138 | 1043 |
| Designer building blocks | 933 | 657 | 617 | 635 |
| Scaffolds | 403 | 353 | 295 | 323 |
| Avg Fsp3 | 0.5 | 0.6 | 0.6 | 0.5 |
| Suitability & Risk | ||||
| Suitability for difficult targets | ★★★ | ★★☆ | ★☆☆ | ★★★ |
| Toxicophores (%) | 1.2 | 1.3 | 1.1 | 2.2 |
| Attachment Chemistry | ||||
| Acylation | ● | ● | ● | ● |
| Reductive amination | ● | ● | ● | ● |
| SNAr | ● | |||
| 1 cyclic tertiary amide (%) | 47 | 50 | 52 | 37 |
| 2 cyclic tertiary amides (%) | 34 | 33 | 57 | |
| Average Physicochemical Properties | ||||
| MW (Da) | 595 | 503 | 525 | 598 |
| cLogP | 2.6 | 0.6 | 0.9 | 1.6 |
| HBA | 8.5 | 6.0 | 6.2 | 6.9 |
| HBD | 2.8 | 2.8 | 2.9 | 2.5 |
| Rotatable bonds | 10.3 | 8.3 | 8.9 | 9.2 |
| TPSA (Ų) | 151 | 135 | 137 | 144 |
Target classes
Our streamlined DNA-encoded chemical library (DEL) screening service for purified proteins has successfully delivered hits and leads against numerous target classes. Including:
Enzymes
- Kinases
- Oxidoreductase Proteases
- Hydrolases
- Isomerases
- Transferases
- Lyases
- Phosphatases
- Transcription factors
- Synthases
- E3 biquitin ligases
- Deubiquitinases (DUBs)
- Helicases
- Polymerases
- Nucleotide exchange factors
- Decarboxylases
- Ligases
Membrane proteins
- Integral membrane proteins
- G protein-coupled receptors (GPCRs)
- Ion channels
- Single-pass membrane proteins
Receptors
- Nuclear receptors
- Cytosolic receptors
- Extracellular receptors
Protein-protein interaction targets
- Transcription factors
- E3 ligases
- Deubiquitinases (DUBs)
- Antibodies
- Cytokines
- Interleukins
- Growth factors
- Carrier proteins
- Adaptor proteins
- Oncoproteins
Epigenetic targets
- Histone binders
- Methyl transferases
- Acetyltransferase
- Demethylases
- Deacetylases
Pathogenic targets
- Viral proteins (enzymes and coat proteins)
- Bacterial proteins (enzymes and receptors)
- Fungal proteins (enzymes and receptors)
Technical information
Related Services
Small molecule drug discovery for even hard-to-drug targets – identify inhibitors, binders and modulators | |
Molecular Glue Direct | |
PPI Inhibitor Direct | |
Integral membrane proteins | |
Selectivity Direct – multiplexed screening of target and anti-targets | |
Express – optimized for fast turn-around-time | |
Snap – easy, fast, and affordable |

