Synthetic Lethality and Context-Dependent Cancer Targeting
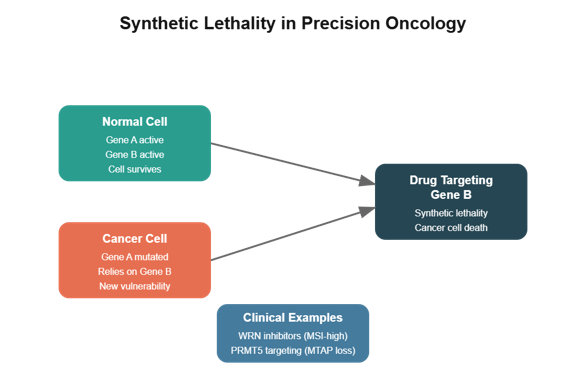
Figure: Conceptual model of synthetic lethality in cancer therapy. In normal cells, redundant pathways allow survival when either gene is inhibited. In cancer cells, mutation of one gene creates dependency on a compensatory pathway. Targeting this remaining pathway produces synthetic lethality, selectively killing tumor cells while sparing normal tissue. Clinically relevant examples include WRN inhibition in MSI-high tumors and PRMT5 targeting in MTAP-deleted cancers.
Executive Overview: From Oncogene Inhibition to Dependency Targeting
For decades, oncology drug discovery focused on inhibiting activated oncogenes — mutant kinases, amplified receptors, dysregulated signaling enzymes. While transformative in many cases, this strategy has intrinsic limits:
- Not all cancers harbor targetable oncogenic mutations
- Many driver proteins lack druggable pockets
- Tumors evolve resistance through pathway rewiring
Synthetic lethality offers a fundamentally different approach. Instead of directly targeting the mutation, it exploits context-specific dependencies created by that mutation.
In simple terms: two genes are synthetically lethal if loss of either alone is tolerated, but loss of both is lethal.
Cancer cells, by virtue of their mutations, often become dependent on backup pathways that normal cells do not require. Synthetic lethality drug discovery aims to identify and therapeutically exploit these vulnerabilities.
The modern era of synthetic lethality encompasses:
- Genome-scale CRISPR screening
- Context-dependent essentiality mapping
- Precision biomarker-driven trials
- Integration with induced proximity therapeutics
Key exemplars include:
- WRN inhibitors in MSI-high cancers
- PRMT5 targeting in MTAP-deleted tumors
- Context-dependent induced proximity strategies
This pillar article explores how synthetic lethality is reshaping precision oncology.
Foundations of Synthetic Lethality
Historical Context
The concept of synthetic lethality originated in classical genetics. In model organisms, researchers observed that deletion of either Gene A or Gene B was tolerated, but deletion of both caused death.
In cancer biology, the idea became clinically validated with PARP inhibitors in BRCA-mutated tumors — demonstrating that therapeutic synthetic lethality could be achieved in patients.
This success shifted oncology strategy from:
- “Target the mutation”
to:
- “Target what the mutation makes essential.”
Feasibility study: 2-4 weeks
Screening: 4-8 weeks
Why Cancer Cells Are Vulnerable
Cancer cells accumulate:
- DNA repair defects
- Replication stress
- Metabolic rewiring
- Proteotoxic stress
- Chromosomal instability
These alterations create dependencies on compensatory pathways. While normal cells retain redundancy, tumor cells often operate at the edge of viability.
Synthetic lethality drug discovery aims to identify these edge conditions.
Precision Oncology Vulnerabilities: Mapping Contextual Dependencies
Modern synthetic lethality relies heavily on:
- CRISPR knockout screens
- RNAi dependency maps
- Multi-omic profiling
- Patient-derived models
Large-scale datasets now catalog gene essentiality across hundreds of cancer cell lines. These maps reveal:
- Lineage-specific vulnerabilities
- Mutation-specific dependencies
- Epigenetic context effects
The challenge lies in translating these genetic interactions into druggable targets with sufficient therapeutic windows.
WRN Dependency in MSI-High Tumors
One of the most compelling recent examples of synthetic lethality involves WRN helicase dependency in microsatellite instability-high (MSI-high) cancers. WRN plays a critical role in DNA replication and repair.
Biological Context: MSI-High Tumors
MSI-high cancers arise from mismatch repair (MMR) deficiency. Loss of MMR leads to:
- Accumulation of insertion/deletion mutations
- Replication stress
- DNA secondary structure instability
To survive this genomic instability, MSI-high tumor cells become dependent on WRN helicase activity.
Mechanistic Basis of WRN Synthetic Lethality
WRN helps resolve:
- DNA secondary structures
- Replication fork stalling
- Microsatellite-induced instability
In MMR-deficient cells, unresolved structures accumulate rapidly without WRN, leading to:
- Chromosomal breakage
- Mitotic catastrophe
- Cell death
In contrast, normal MMR-proficient cells tolerate WRN inhibition more effectively.
Therapeutic Development
WRN inhibitors represent a promising strategy in MSI-high colorectal, gastric, and endometrial cancers.
Advantages include:
- Strong genetic validation
- Biomarker clarity (MSI-high status)
- Potential synergy with immunotherapy
Challenges remain:
- Achieving selective helicase inhibition
- Avoiding toxicity in proliferative tissues
- Managing resistance through compensatory repair pathways
WRN inhibition exemplifies synthetic lethality driven by DNA repair deficiency.
PRMT5 Dependency in MTAP-Deleted Tumors
Another well-characterized vulnerability involves PRMT5 inhibition in tumors with MTAP deletion. PRMT5 is an epigenetic regulator involved in symmetric arginine methylation. MTAP is frequently co-deleted with CDKN2A in many cancers.
Biological Context
MTAP deletion leads to accumulation of methylthioadenosine (MTA), a metabolite that partially inhibits PRMT5. This creates a state where:
- PRMT5 activity is already compromised
- Cancer cells operate near minimal PRMT5 function
- Further inhibition becomes selectively lethal
Mechanistic Insight
PRMT5 regulates:
- Splicing
- Chromatin modification
- Transcriptional control
MTAP-deleted cells are hypersensitive to PRMT5 inhibition because MTA accumulation lowers the functional threshold.
Therapeutic approaches include:
- Substrate-competitive inhibitors
- MTA-cooperative inhibitors
- Context-selective PRMT5 targeting
Clinical Implications
MTAP deletion is common across:
- Glioblastoma
- Pancreatic cancer
- Lung cancer
- Mesothelioma
This provides a broad patient population with defined biomarker stratification.
Key development challenges:
- Avoiding hematologic toxicity
- Optimizing therapeutic index
- Understanding splicing-related side effects
PRMT5–MTAP represents a metabolic-epigenetic synthetic lethal axis.
Induced Proximity and Context-Dependent Targeting
Beyond classical enzyme inhibition, induced proximity technologies are beginning to intersect with synthetic lethality strategies. RIPTACs represent one such example.
Context-Dependent Induced Proximity
RIPTACs force interaction between:
- A broadly expressed essential protein
- A tumor-restricted protein
The essential protein becomes functionally impaired only in cells expressing the tumor-specific anchor. This creates synthetic lethality without directly targeting a mutated gene.
Advantages
- Expanded druggable space
- Selectivity driven by co-expression
- Potential targeting of non-mutant tumors
Induced proximity adds a programmable dimension to synthetic lethality.
Discovery Strategies in Synthetic Lethality Drug Discovery
CRISPR-Based Screens
Genome-wide CRISPR knockout or CRISPR interference screens identify:
- Mutation-specific essential genes
- Pathway dependencies
- Synergistic vulnerabilities
High-throughput combinatorial screens accelerate mapping of lethal gene pairs.
Multi-Omic Integration
Synthetic lethality is rarely binary. Context matters:
- Tumor lineage
- Epigenetic state
- Metabolic environment
- Microenvironmental stress
Integrating genomics, transcriptomics, proteomics, and metabolomics improves predictive accuracy.
Patient Stratification
Precision oncology vulnerabilities require robust biomarkers:
- MSI-high status for WRN
- MTAP deletion for PRMT5
- Specific mutation signatures
Without clear stratification, therapeutic windows collapse.
Resistance Mechanisms
As with any targeted therapy, resistance can emerge. Common resistance pathways include:
- Restoration of lost repair functions
- Upregulation of compensatory pathways
- Mutation of drug target
- Epigenetic adaptation
For example:
- Alternative helicases may compensate for WRN inhibition
- PRMT5 pathway rewiring may restore splicing balance
Combination strategies are often necessary to sustain efficacy.
Combination Strategies
Synthetic lethality often synergizes with:
- Immunotherapy
- DNA-damaging agents
- Targeted protein degradation
- Epigenetic modulators
For example:
- WRN inhibition may increase tumor mutational burden and immune visibility
- PRMT5 inhibition may sensitize tumors to splicing stress
Rational combinations are critical for durable response.
Expanding the Synthetic Lethal Landscape
Emerging directions include:
- Targeting replication stress pathways
- Exploiting metabolic bottlenecks
- Synthetic lethal interactions with chromatin remodelers
- Leveraging tumor microenvironment dependencies
AI-driven models may soon predict synthetic lethal interactions de novo.
Commercial and Translational Landscape
Synthetic lethality drug discovery has become a cornerstone of precision oncology pipelines.
Pharmaceutical interest is high because:
- Genetic validation reduces target risk
- Biomarker-defined populations improve trial efficiency
- First-in-class opportunities are abundant
Companies are investing in:
- Dependency mapping platforms
- CRISPR-enabled discovery
- Computational vulnerability prediction
Synthetic lethality is now integrated into mainstream oncology R&D strategy.
Future Outlook: From Mutation-Driven to Dependency-Driven Oncology
The future of cancer therapy may depend less on targeting oncogenes directly and more on targeting what cancer cells cannot live without.
Synthetic lethality reframes cancer not as a disease of activation alone, but as a disease of vulnerability.
Key future developments may include:
- Personalized dependency maps
- Dynamic vulnerability profiling during treatment
- Synthetic lethal networks rather than single pairs
- Integration with spatial and proximity-based therapeutics
Precision oncology vulnerabilities are likely to become increasingly refined as tumor heterogeneity is better understood.
Conclusion
Synthetic lethality drug discovery represents one of the most rational and genetically grounded approaches in modern oncology.
From WRN inhibitors in MSI-high cancers to PRMT5 targeting in MTAP-deleted tumors, context-specific vulnerabilities are yielding actionable therapeutic strategies.
The core insight is simple but powerful: cancer cells survive by rewiring biology — and that rewiring creates dependencies.
By systematically identifying and exploiting those dependencies, precision oncology moves beyond inhibiting drivers to dismantling survival scaffolds.
As discovery technologies mature and induced proximity tools expand the targetable space, synthetic lethality will continue to redefine how cancer is treated — not by attacking what tumors are, but by exploiting what they cannot live without.
Related Services
| Service | |
|---|---|
Small molecule drug discovery for even hard-to-drug targets – identify inhibitors, binders and modulators | |
Molecular Glue Direct | |
PPI Inhibitor Direct | |
Integral membrane proteins | |
Specificity Direct – multiplexed screening of target and anti-targets | |
Express – optimized for fast turn – around-time | |
Snap – easy, fast, and affordable |


