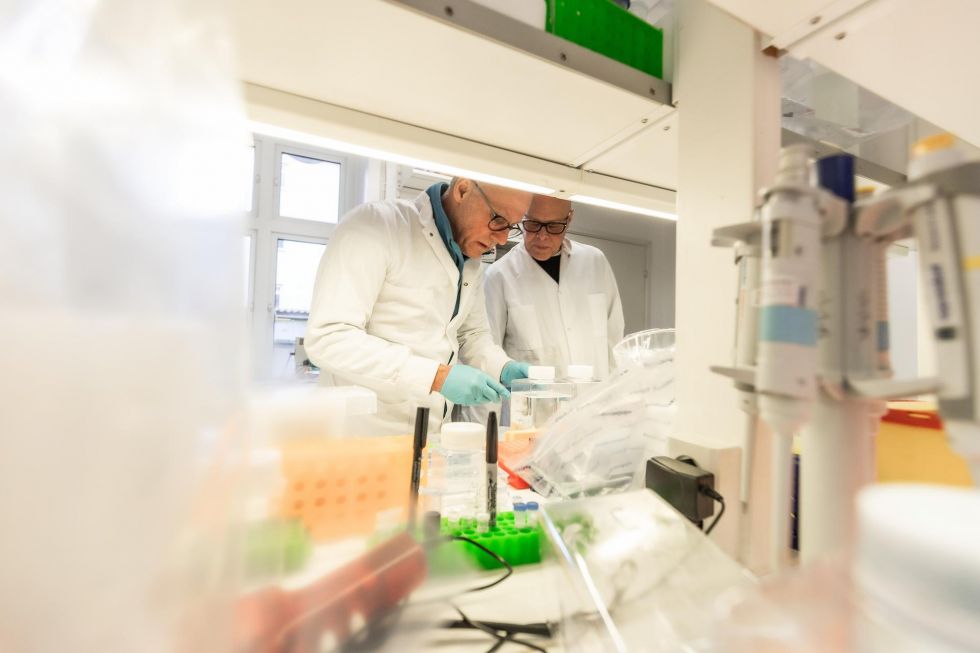
Autophagy and Lysosome-Based Targeted Degradation Technologies
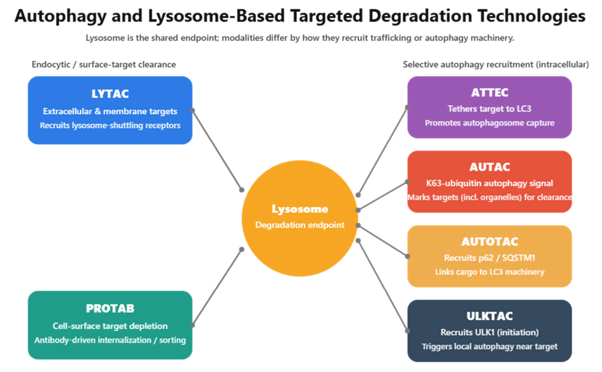
Figure: Overview of lysosome- and autophagy-based targeted degradation technologies.
Lysosomal degradation pathways expand targeted protein degradation beyond the ubiquitin–proteasome system. LYTACs and PROTABs enable clearance of extracellular and membrane proteins through receptor-mediated endocytosis and lysosomal trafficking. Intracellular autophagy-based systems include ATTECs (LC3 tethering), AUTACs (K63-linked ubiquitin autophagy signaling), AUTOTACs (p62-mediated recruitment), and ULKTACs, which initiate autophagy through ULK1 activation. Together these technologies broaden the degradable target space to include extracellular proteins, aggregates, and organelles.
Beyond the Proteasome: Why Lysosomal Pathways Matter
Targeted protein degradation (TPD) initially centered on the ubiquitin–proteasome system (UPS). Classical PROTACs demonstrated that small molecules could catalytically eliminate intracellular proteins by recruiting E3 ligases. Yet the proteasome has intrinsic constraints:
- It primarily degrades soluble intracellular proteins
- It struggles with large aggregates and organelles
- It cannot directly eliminate extracellular or membrane proteins
- Its activity can be compromised in stressed or diseased cells
These limitations have catalyzed the rise of lysosome- and autophagy-based targeted degradation technologies — modalities that harness the cell’s broader proteostasis and trafficking machinery.
Unlike the proteasome, the lysosome:
- Degrades membrane proteins and extracellular ligands
- Handles protein aggregates and damaged organelles
- Integrates with endocytosis and macroautophagy
- Provides access to previously “undruggable” spatial compartments
This expansion of degradable target space is reshaping therapeutic strategies in oncology, neurodegeneration, immunology, and rare disease.
Lysosome-Targeting Chimeras (LYTACs): Clearing the Extracellular Proteome
Mechanistic Overview
LYTACs are bifunctional molecules that bind:
- An extracellular or membrane-bound target protein
- A lysosome-shuttling receptor (such as CI-M6PR or ASGPR)
Upon binding, the ternary complex is internalized through endocytosis and trafficked to the lysosome for degradation.
This enables:
- Degradation of secreted cytokines
- Removal of membrane receptors
- Clearance of pathogenic extracellular proteins
Unlike monoclonal antibodies, which typically block function, LYTACs eliminate the target entirely.
Receptor Systems Used in LYTAC Design
CI-M6PR (Cation-Independent Mannose-6-Phosphate Receptor)
- Broad tissue expression
- Established lysosomal trafficking route
- Suitable for systemic applications
ASGPR (Asialoglycoprotein Receptor)
- Liver-restricted expression
- Enables hepatocyte-specific degradation
- Valuable for metabolic and liver diseases
Receptor selection determines tissue specificity and therapeutic index.
Therapeutic Applications of LYTACs
Oncology
- Degradation of growth factor receptors
- Removal of immune checkpoint ligands
- Targeting shed extracellular drivers
Immunology
- Cytokine clearance
- Modulation of inflammatory mediators
Fibrotic Disease
- Eliminating profibrotic secreted proteins
LYTACs extend TPD into compartments previously inaccessible to small molecules.
LYMTACs and PROTABs: Next-Generation Surface Target Clearance
Beyond classical LYTACs, newer modalities refine surface protein degradation strategies.
LYMTACs (Lysosomal Membrane Targeting Chimeras)
LYMTACs expand lysosome-directed degradation by targeting lysosomal membrane proteins or alternative internalization routes.
Key differentiators:
- Exploit distinct lysosomal membrane trafficking pathways
- Potentially bypass limitations of CI-M6PR dependency
- Broaden receptor options
LYMTACs increase flexibility in receptor biology and tissue targeting.
PROTABs (Proteolysis-Targeting Antibodies)
PROTABs are antibody-based constructs designed to induce degradation of cell-surface targets.
Mechanistic logic:
- Antibody binds membrane target
- Engages endogenous ubiquitination or internalization machinery
- Drives lysosomal degradation
Unlike classical antibodies that merely block receptor function, PROTABs induce complete target removal.
Advantages over mAbs:
- Sustained target depletion
- Reduced compensatory signaling
- Potential to overcome resistance to receptor blockade
Selective Autophagy-Based Targeted Degradation
While LYTACs operate via endocytic–lysosomal pathways, other technologies harness macroautophagy, the cell’s bulk degradation system.
Autophagy normally clears:
- Protein aggregates
- Damaged mitochondria
- Intracellular pathogens
Selective autophagy can be induced through engineered proximity strategies.
ATTECs: Autophagy-Tethering Compounds
ATTECs are small molecules that bind both:
- A target protein
- LC3 (a core autophagosome protein)
By tethering the target to LC3, ATTECs promote selective autophagic engulfment and degradation.
Key Advantages
- Proteasome-independent
- Effective against aggregate-prone proteins
- Promising in neurodegenerative diseases
Limitations
- Dependence on autophagic flux
- Potential off-target autophagy activation
AUTACs: Autophagy-Targeting Chimeras
AUTACs function differently from ATTECs.
Mechanism
AUTACs mimic S-guanylation signals that induce K63-linked ubiquitination. This modification marks the target for selective autophagic degradation.
Distinguishing Features
- Ubiquitin-dependent autophagy engagement
- Broader compatibility with cellular degradation signals
- Potential organelle targeting applications
AUTACs have been explored for mitochondrial clearance and damaged organelle removal.
AUTOTACs: p62-Mediated Selective Autophagy
AUTOTACs directly recruit the autophagy receptor p62/SQSTM1 to a target.
Mechanistic Insight
p62 recognizes ubiquitinated proteins and links them to LC3 on autophagosomes. AUTOTACs bypass ubiquitination complexity by directly engaging p62.
Advantages
- Simplified recruitment logic
- Potentially more direct autophagy initiation
ULKTACs: Initiating Autophagy at the Source
ULKTACs recruit ULK1, the kinase that initiates autophagosome formation.
Instead of tagging the target for existing autophagy, ULKTACs locally activate autophagy machinery near the target protein.
Implications
- Spatially confined autophagy
- Greater control of degradation initiation
- Potential to overcome autophagic flux limitations
Comparing Autophagy and Lysosome Modalities
| Feature | LYTAC | ATTEC | AUTAC | AUTOTAC | ULKTAC |
|---|---|---|---|---|---|
| Target location | Extracellular/membrane | Intracellular | Intracellular | Intracellular | Intracellular |
| Ubiquitin required | No | No | Yes | Often | Variable |
| Receptor dependency | Yes | LC3 | K63 ubiquitin | p62 | ULK1 |
| Organelles targetable | Limited | Possible | Yes | Possible | Potentially broad |
Design and Discovery Considerations
Autophagic Flux Competence
Many diseases (e.g., neurodegeneration) involve impaired autophagy. Therapeutic success depends on:
- Baseline flux levels
- Lysosomal integrity
- mTOR signaling state
Biomarker development is essential.
Tissue Selectivity
Lysosome and autophagy components vary across tissues. Leveraging:
- Receptor-restricted expression (ASGPR)
- Tissue-specific autophagy regulators
- Delivery vectors
can optimize therapeutic index.
Avoiding Global Autophagy Activation
Non-selective autophagy induction risks:
- Muscle wasting
- Immune dysregulation
- Metabolic disturbance
Precision recruitment is therefore critical.
Structural and Chemical Design
Challenges include:
- Large molecular weight
- Cell permeability
- Endosomal escape (for extracellular targeting systems)
- Maintaining ternary complex stability
Structure-guided design and high-content imaging screens are increasingly central.
Disease Applications
Neurodegeneration
Autophagy-based degraders are especially promising in:
- Aggregate-prone protein diseases
- Huntington’s disease
- Tauopathies
- Parkinson’s disease
Proteasome systems often struggle with large fibrillar aggregates; autophagy excels in this domain.
Oncology
In cancer, autophagy plays dual roles:
- Tumor suppression (via proteostasis maintenance)
- Tumor survival under stress
Targeted autophagic degradation can:
- Eliminate oncogenic drivers
- Remove immune evasion proteins
- Target metabolic dependencies
Immunology and Inflammation
Selective clearance of:
- Cytokines
- Surface immune checkpoints
- Secreted inflammatory mediators
could reshape autoimmune and inflammatory disease treatment.
Rare Genetic Disorders
Diseases caused by:
- Toxic gain-of-function proteins
- Organelle accumulation
- Protein misfolding
may benefit from selective autophagy recruitment strategies.
Translational and Competitive Landscape
Biotech companies are rapidly expanding into lysosomal and autophagy platforms, often positioning themselves as:
- “Beyond PROTAC” companies
- Extracellular degradome specialists
- Autophagy-first drug discovery platforms
Key differentiators include:
- Receptor libraries
- Autophagy receptor selectivity
- Tissue-targeting technologies
- Biomarker development capabilities
Pharmaceutical interest is growing due to:
- Access to membrane proteins
- Potential first-in-class opportunities
- Synergy with antibody platforms
Resistance and Adaptive Responses
As with proteasomal degradation, resistance can emerge through:
- Receptor downregulation
- Lysosomal dysfunction
- Autophagy pathway mutations
- Compensatory signaling pathways
Combination strategies (e.g., mTOR modulation + targeted autophagy) may improve durability.
The Future of Lysosome and Autophagy-Based Pharmacology
The next phase of innovation will likely focus on:
- Organelle-specific degradation (mitophagy-targeting chimeras)
- Condensate-associated autophagy
- Tissue-restricted lysosomal receptors
- Small-molecule modulators of autophagy selectivity
- Multi-functional chimeras integrating degradation and signaling control
Integration with:
- Synthetic lethality frameworks
- Spatial pharmacology
- AI-guided ternary complex modeling
will further refine precision.
Conclusion: Expanding the Degradable Universe
Autophagy- and lysosome-based targeted degradation technologies represent a decisive expansion of the druggable proteome.
They enable:
- Extracellular protein elimination
- Aggregate clearance
- Organelle removal
- Context-dependent degradation
Where classical PROTACs opened the door to catalytic protein elimination, lysosomal and autophagy modalities unlock the broader spatial architecture of cellular proteostasis.
The field is evolving from simply degrading intracellular proteins to programming cellular disposal systems.
In doing so, drug discovery is transitioning from blocking disease drivers to orchestrating their controlled removal across compartments — inside, outside, and across organelles.
The proteasome was just the beginning.
Related Services
| Service | |
|---|---|
Small molecule drug discovery for even hard-to-drug targets – identify inhibitors, binders and modulators | |
Molecular Glue Direct | |
PPI Inhibitor Direct | |
Integral membrane proteins | |
Specificity Direct – multiplexed screening of target and anti-targets | |
Express – optimized for fast turn – around-time | |
Snap – easy, fast, and affordable |


