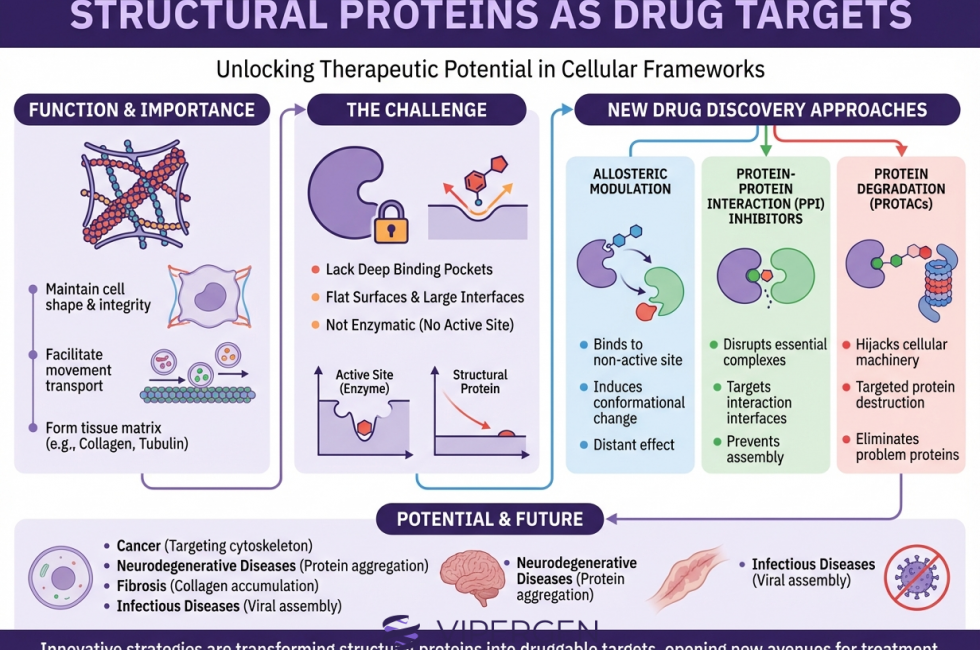
Structural Proteins as Drug Targets: In-Cell DEL Screening for Oncology Programs
Structural proteins sit at the heart of cancer cell biology. They build the cytoskeleton (microtubules, actin filaments, intermediate filaments), organize the mitotic spindle, shape membranes, and coordinate transport, polarity, invasion in cancer, and survival. Because these systems are essential for proliferation and metastasis, they have also produced some of the most durable drug classes in oncology, especially microtubule-targeting agents used across solid tumors and hematological malignancies [1], [2].
At the same time, “structural protein drug discovery” has historically come with tradeoffs: narrow therapeutic windows, complex cellular context, and targets that are dynamic, polymeric, and highly conserved. These challenges are why many cytoskeletal drug targets still rely on legacy chemotypes or phenotypic screening hits, rather than modern, binding-driven campaigns. Yet the last decade has brought a major shift in what is possible: improved structural biology (cryo-EM of polymeric complexes), better in-cell target engagement assays, and DNA-encoded library (DEL) methods that can generate binders for hard-to-drug proteins, including in living cells.
This article maps the opportunity landscape for cytoskeleton drug targets (with emphasis on microtubule ligand discovery and antimitotic programs), outlines why structural proteins are “binding-driven targets” despite their reputation, and explains how DNA encoded library oncology screening (including in-cell approaches) can complement (and de-risk) traditional phenotypic screening.
Structural Proteins in Cellular Architecture
Structural proteins are often described as “scaffolding,” but in cancer biology they are better understood as adaptive machines. They remodel in seconds, respond to force, and integrate signaling and metabolism. That dynamism is precisely what makes them attractive, albeit difficult drug targets.
Cytoskeletal Dynamics
The cytoskeleton is not one system; it is a coordinated network:
- Microtubules: polarized polymers of α/β-tubulin that form tracks for kinesin/dynein transport, organize organelles, and build the mitotic spindle.
- Actin filaments: rapidly assembling networks that generate force for migration, cytokinesis, and membrane remodeling.
- Intermediate filaments: stress-bearing polymers (e.g., vimentin, keratins) that provide mechanical resilience and influence signaling states.
Crosstalk between actin and microtubules is central to migration and division and provides multiple intervention points beyond “direct tubulin binders” [3]. Natural products have historically been a key source of cytoskeletal probes and drugs, highlighting that these proteins are chemically addressable even when they appear “flat” in simplified models [4].
Role in Cell Division and Survival
Cytoskeletal proteins become most targetable in oncology when one connects them to vulnerabilities:
- Mitosis and chromosomal instability: spindle assembly, microtubule dynamics, and checkpoint timing are sensitive to small changes in polymer behavior – one reason antimitotic drug discovery continues to evolve [1], [2].
- Metastasis and invasion: actin remodeling, adhesion turnover, and intermediate filament switching (e.g., vimentin upregulation) are connected to Epithelial–Mesenchymal Transition (EMT)-like programs and aggressive phenotypes [3], [7].
- Stress tolerance: intermediate filaments buffer mechanical stress and influence organelle positioning; their dysregulation links to human disease and cancer biology [6].
The key takeaway for drug hunters: structural proteins are essential, but their essentiality is conditional; cell type, state, microenvironment, and genetic background matter. That is both the risk and the opportunity.
Structural Protein Targets Accessible in Cells
When people say, “structural proteins,” they often mean tubulin and actin. In practice, oncology-relevant “structural protein targets” span: polymers (tubulin/actin), polymer regulators, spindle machinery, and cytoskeleton-associated complexes.
Microtubules
Microtubules remain the flagship example of a structural protein drug target class. Multiple binding sites on tubulin enable stabilizers and destabilizers, and clinically validated mechanisms include both mitotic arrest and non-mitotic effects on trafficking, signaling, and tumor vasculature [1], [2].
What’s changing now is not the importance of tubulin, but the precision with which we can define where and how a ligand binds:
- Structural methods have mapped canonical sites (taxane, vinca alkaloids, colchicine, etc. [Scheme 1]) and revealed how distinct ligands tune microtubule dynamics [22]–[24].
- The “tubulin code” – isotypes and post-translational modifications – adds layers of biological selectivity that can be exploited for context-specific targeting and safety [21].
Scheme 1: Structure of different tubulin ligands.
From program-design perspective, this is where microtubule ligand discovery becomes more than “find another tubulin poison.” The frontier is selective modulation: isoform bias, complex-specific binding, or allosteric control of partner proteins.
Mitotic Spindle Components
“Antimitotic drug discovery” is broader than tubulin. Many programs target spindle machinery indirectly:
- Motor proteins (e.g., kinesins such as Eg5/KIF11)
- Microtubule-associated proteins (MAPs)
- Spindle assembly and checkpoint regulators
- Protein–protein interactions (PPIs) within dynamic mitotic complexes
A classic example is the discovery of monastrol, a small molecule that perturbs spindle bipolarity by inhibiting the kinesin Eg5 – an early demonstration that mitotic machinery can yield tractable small-molecule binders even outside tubulin [25].
Beyond Tubulin: Actin and Intermediate Filaments
Actin is deeply implicated in invasion and survival, but direct actin targeting has been limited clinically by toxicity. A major strategy is to target actin regulators (e.g., tropomyosins and nucleation factors), rather than the core polymer [5]. Intermediate filaments are less explored but increasingly recognized as druggable via indirect mechanisms (disrupting assembly, targeting associated kinases, or exploiting cancer-specific dependencies). Their disease links and mechanistic roles are well established [6], and vimentin’s role in cancer phenotypes makes it a frequent point of interest [7].
Drug Discovery Challenges
Structural proteins generate three recurring challenges that directly impact hit discovery and lead optimization.
Balancing Potency and Cellular Toxicity
For essential cellular machines, potency is easy to achieve and hard to use safely. This is one reason microtubule drugs can be effective yet limited by neuropathy or myelosuppression, and why actin-directed approaches can be constrained by cardio-toxicity risks [2], [5].
Practical implications for discovery teams:
- Avoiding “global polymer poisons”: prioritize modulators that bias specific states, complexes, or cell contexts.
- Separating binding from catastrophic function loss: not every binder must be a strong destabilizer; some can be allosteric modulators or complex stabilizers.
Early safety triage: structural proteins demand early counterscreens and cellular selectivity analysis, not late-stage “surprises.”
Context-Specific Target Engagement
Structural proteins are dynamic and compartmentalized. The “same” protein can exist in multiple functional forms:
- Soluble monomer/dimer pools vs polymerized filaments
- Distinct complexes with partner proteins
- Isoforms and post-translational modification (PTM)-defined subpopulations (notably for tubulin) [21]
That means a binder found against purified protein may not engage the relevant cellular state. This is where protein binding site identification and in-cell target engagement become central – not optional.
Two widely used families of approaches are:
- CETSA (Cellular Thermal Shift Assay) to measure stabilization of proteins upon ligand binding in cells/tissues [19]
- Thermal proteome profiling (TPP) to assess proteome-wide engagement and downstream effects in living cells [20]
These methods don’t replace functional assays, but they reduce ambiguity early especially when structural proteins generate complex phenotypes.
Protein Binding Site Identification for Structural Proteins
Why “Structural” Doesn’t Mean “Undruggable”
A misconception is that structural proteins lack binding pockets. In reality, they often offer:
- Interfacial pockets (between subunits or protein partners)
- Conformation-specific cavities (open only in certain states)
- Polymer-specific surfaces (present only in filaments)
- Allosteric sites that tune dynamics rather than block an active site
Tubulin illustrates this best: decades of work have mapped major small-molecule sites and explained how different ligands stabilize or destabilize microtubules [22]–[24]. The emerging frontier is expanding beyond “the usual sites” and leveraging complex-dependent pockets.
A recent example: a newly defined tubulin ligand-binding site
A compelling recent study reported a previously unknown tubulin ligand-binding site that only emerges in the context of a tubulin complex with the stathmin family protein RB3 and tubulin tyrosine ligase (TTL) [26]. The authors show that Tumabulin-1 (TM1, Scheme 2), derived from BML284, can bind not only the canonical colchicine site but also a second site located at an interface involving α/β-tubulin plus RB3. Strikingly, two TM1 molecules bind cooperatively in this relatively large pocket, contacting multiple proteins in the complex. The work goes further by designing Tumabulin-2 (TM2, Scheme 2), a derivative engineered to bind selectively to this newly described “Tumabulin site” rather than the colchicine site. Functionally, TM2 behaves as a molecular glue: it strengthens the interaction between RB3 and tubulin, enhances RB3’s tubulin-depolymerizing activity, and supports the idea that “complex-first” pockets can be exploited to create new antimitotic mechanisms. Conceptually, this is a blueprint for future microtubule ligand discovery: instead of only targeting tubulin in isolation, drug discovery can aim at partner-dependent binding sites that may offer more selective biology and new safety/efficacy tradeoffs.
Scheme 2: Structure of TM1 and TM2
Structural Proteins as Binding-Driven Targets
Structural proteins are often discovered via phenotypic effects (mitotic arrest, migration block). But that doesn’t mean they are “phenotype-only” targets. They are profoundly binding-driven:
- A small change in microtubule catastrophe frequency can shift mitotic outcomes [1], [2]
- Stabilizing or destabilizing specific actin architectures can alter cancer invasion programs [3], [5]
- Modulating complex formation (molecular glue mechanisms) can create new functional outputs [26]
This is the bridge to DEL: DEL screening is best when binding is the primary signal and when rapid exploration of chemical space is wanted to find rare chemotypes that can engage challenging pockets [11], [12].
In-Cell DEL Screening for Structural Proteins
Why in-cell approaches matter for cytoskeleton drug targets
Structural proteins are context-dependent; many relevant binding sites are:
- conformation-specific,
- complex-dependent,
- influenced by PTMs/isoforms,
- or only present in living-cell conditions.
In-cell approaches reduce the risk of selecting binders that only recognize an artificial purified state.
What DEL is (and why it’s relevant here)
DNA-encoded libraries link each small molecule to a DNA barcode so billions of compounds can be pooled, selected for binding, and decoded by sequencing. The foundational concept of encoding chemistry with amplifiable information dates back to early encoded combinatorial chemistry [10], and has since matured into a widely used hit-finding engine [11]. Modern reviews cover selection formats, library design, artifacts, and translation to medicinal chemistry [12], [13].
Evidence base for DEL screening on or in living cells
Several peer-reviewed studies established that DEL selections can be done in living-cell contexts, including on-cell-surface targets and intracellular environments:
- DEL selection within and on living cells using cell-penetrating approaches [14]
- DEL screening inside a living cell using Xenopus laevis oocytes as a cellular “reaction vessel” [15]
- DEL selection against endogenous membrane proteins on live cells, supporting native-context binder discovery without purified target [16]
For structural proteins and cytoskeleton-associated complexes, these approaches are attractive because they better preserve polymer states, native assemblies, and cellular competition effects – key determinants of “real” engagement.
DEL screening directly in living cells
Vipergen has described service and technology approaches that emphasize DEL screening from target binding through hit validation, including screening formats designed for living-cell contexts and difficult targets [17]. Vipergen also offers a “Molecular Glue Direct” DELs-in-cells format intended to discover molecular glues by co-expressing targets and interaction partners (including E3 ligases) and screening in a multiplexed format in living cells [18]. For structural proteins, this matters because complex-dependent pockets (like the Tumabulin site example above) are often the most compelling places to look – but they are also the easiest to miss in simplified assays.
DEL Workflow and Applications for Oncology Programs
1) Define the structural target hypothesis (binder-first)
For cytoskeleton drug targets, the hypothesis is usually one of these:
- Polymer modulation: tune microtubule/actin dynamics without total collapse
- Complex modulation: target a partner-dependent interface pocket (e.g., tubulin-RB3-like concepts)
- Spindle machinery inhibition: block a motor protein or mitotic complex assembly
- Selectivity-by-context: exploit isoforms/PTMs/complexes that differ between tumor and normal tissues [21]
In practical terms: The question is not only asking “can we inhibit this?” but “which binding event produces the oncology-relevant phenotype with acceptable safety?”
2) Choose the selection environment: in vitro vs in-cell
- In vitro DEL screening can be ideal when stable purified protein is easily accessible, tight control is wanted and can fold into the relevant conformation.
- In-cell DEL screening becomes especially valuable when:
- The target is hard to purify
- The relevant binding site is complex-dependent
- Post-translational modifications matter,
- Early evidence of cellular relevance is wanted [14-16].
For cytoskeleton targets, in-cell approaches can also help filter out compounds that bind the purified protein but cannot compete with endogenous binding partners or fail due to localization barriers.
3) Run a selection designed to reduce “structural protein artifacts”
Structural proteins are abundant and sticky; selections must be designed to reduce frequent hitters and nonspecific binders. Typical design principles include:
- Multiple negative selections (beads/DNA controls; off-target protein controls)
- Competition experiments with known ligands (when available)
- Orthogonal selection conditions (salt, detergents compatible with target state, polymer vs monomer conditions)
- Early clustering and “chemical series” prioritization rather than singletons
The point is not to eliminate all noise, it’s to quickly surface chemotypes that behave consistently across selection variants.
4) Translate hits off-DNA and validate binding
DEL delivers hypotheses; medicinal chemistry needs confirmed molecules. Typical validation includes:
- Biophysical binding confirmation (MST, SPR, ITC, DSF)
- Structural follow-up (cryo-EM or X-ray where feasible), particularly important for tubulin sites [22-24]
- Cellular target engagement:
- CETSA to confirm in-cell binding [19]
- TPP to assess proteome-wide engagement and potential liabilities [20]
For structural proteins, engagement data is especially useful because phenotypes can be indirect. Binding confirmation quickly identifies wrong mechanism which can then be eliminated.
5) Functional assays: tune the phenotype, not just potency
For oncology, the functional readouts depend on target class:
- Microtubules: polymerization dynamics, spindle formation, mitotic timing, aneuploidy signatures
- Spindle motors: bipolar spindle assembly defects, mitotic arrest phenotypes (monastrol-like patterns) [25]
- Actin systems: migration/invasion assays, adhesion turnover, cytokinesis defects [3], [5]
- Complex-dependent targets: specific biomarker readouts tied to complex stabilization/destabilization (as in molecular glue concepts) [26]
A practical strategy is to start with a binder series, then intentionally explore the “phenotype space” by tuning permeability, residence time, and allosteric bias – rather than pushing raw potency immediately.
Positioning Against Phenotypic Screening
Phenotypic screening has historically been productive in cytoskeletal biology – partly because many phenotypes (mitotic arrest, rounded morphology, migration block) are easy to observe. But phenotypic screening also has recurring major bottlenecks and constraints:
- target deconvolution can be slow,
- the same phenotype can come from many targets,
- and off-target toxicity can generate false-positive signals / artifactual activity as “activity.”
Large analyses of discovery strategies have shown that phenotypic approaches have contributed substantially to first-in-class medicines [8], and industry perspectives continue to highlight both opportunity and friction in phenotypic drug discovery [9].
DEL-based discovery offers a complementary angle:
- DEL is binder-first: Chemical matter is the starting point which can be tied to a target hypothesis.
- Phenotypic screens are effect-first: Phenotype is the starting point and search for mechanism is the later work.
For structural proteins, a hybrid approach is often best:
- use binding-driven DEL screening to generate selective chemotypes for a structural target or complex,
- then test those chemotypes across phenotypic systems (cell division, migration, survival),
- while using target engagement tools (CETSA/TPP) to confirm that cellular biology matches the binding hypothesis [19], [20].
This is especially valuable for modern microtubule ligand discovery, where the goal is not “maximum disruption,” but mechanism-shaped modulation that delivers therapeutic windows.
Practical “High-Intent” Use Cases in Oncology
To align with real search behavior and program decisions, here are common scenarios where DEL screening for structural protein targets is often considered:
Use case A: Replace or differentiate legacy microtubule chemotypes
A team wants microtubule modulation but needs:
- a new binding site,
- less neuropathy risk,
- a differentiated resistance profile,
- or a complex-dependent mechanism.
The Tumabulin site example shows how new tubulin pockets can emerge in complexes and enable novel mechanisms like molecular glue-like stabilization of protein–protein interactions [26].
Use case B: Move from “antimitotic phenotype” to a defined mitotic target
Phenotypic screens find mitotic arrest hits, but target IDs are unclear. DEL can support a binder-first campaign against a mitotic motor or spindle complex, inspired by precedents like Eg5 inhibition [25].
Use case C: Target migration/invasion without direct actin poisoning
Instead of actin itself, target actin regulators or cross-talk nodes that influence cytoskeletal architecture [3], [5]. DEL can help identify small-molecule binders for proteins that are hard to screen traditionally.
Conclusion: Structural Proteins Are Back – With Better Tools
Structural proteins are no longer “legacy-only” oncology targets. Modern structural biology is exposing new pockets in polymeric complexes [22-24] and even expanding the tubulin site landscape through complex-dependent binding sites and molecular glue concepts [26]. In parallel, target engagement methods like CETSA and TPP reduce ambiguity about what a compound is doing in cells [19], [20], and DEL technologies keep expanding the practical chemical search space for hard-to-drug targets [11-13].
For teams working on structural protein drug discovery, the strategic shift is this: move from “find a toxin” to “find a binder that modulates a defined state or complex in the right cellular context.” That’s where DNA encoded library oncology screening – including in-cell formats – can deliver value: fast hit generation, early selectivity signals, and a clearer line from binding to biology [14]–[18].
References
[1] Jordan M.A., Wilson L., Microtubules as a target for anticancer drugs, Nat. Rev. Cancer (2004), 4, 253–265. https://doi.org/10.1038/nrc1317
[2] Dumontet C., Jordan M.A., Microtubule-binding agents: a dynamic field of cancer therapeutics, Nat. Rev. Drug Discov. (2010), 9, 790–803. https://doi.org/10.1038/nrd3253
[3] Dogterom M., Koenderink G.H., Actin–microtubule crosstalk in cell biology, Nat. Rev. Mol. Cell Biol. (2019), 20, 38–54. https://doi.org/10.1038/s41580-018-0067-1
[4] Risinger A.L., Du L., Targeting and extending the eukaryotic druggable genome with natural products: cytoskeletal targets of natural products, Nat. Prod. Rep. (2020), 37, 634–652. https://doi.org/10.1039/C9NP00053D
[5] Bryce N.S., Hardeman E.C., Gunning P.W., Lock J.G., Chemical biology approaches targeting the actin cytoskeleton through phenotypic screening, Curr. Opin. Chem. Biol. (2019), 51, 40–47. https://doi.org/10.1016/j.cbpa.2019.02.013
[6] Omary M.B., Coulombe P.A., McLean W.H.I., Intermediate filament proteins and their associated diseases, N. Engl. J. Med. (2004), 351, 2087–2100. https://doi.org/10.1056/NEJMra040319
[7] Satelli A., Li S., Vimentin in cancer and its potential as a molecular target for cancer therapy, Cell. Mol. Life Sci. (2011), 68, 3033–3046. https://doi.org/10.1007/s00018-011-0735-1
[8] Swinney D.C., Anthony J., How were new medicines discovered?, Nat. Rev. Drug Discov. (2011), 10, 507–519. https://doi.org/10.1038/nrd3480
[9] Moffat J.G., Vincent F., Lee J.A., et al., Opportunities and challenges in phenotypic drug discovery: an industry perspective, Nat. Rev. Drug Discov. (2017), 16, 531–543. https://doi.org/10.1038/nrd.2017.111
[10] Brenner S., Lerner R.A., Encoded combinatorial chemistry, Proc. Natl. Acad. Sci. U.S.A. (1992), 89(12), 5381–5383. https://doi.org/10.1073/pnas.89.12.5381
[11] Goodnow R.A. Jr., Dumelin C.E., Keefe A.D., DNA-encoded chemistry: enabling the deeper sampling of chemical space, Nat. Rev. Drug Discov. (2017), 16, 131–147. https://doi.org/10.1038/nrd.2016.213
[12] Neri D., Lerner R.A., DNA-encoded chemical libraries: a selection system based on endowing organic compounds with amplifiable information, Annu. Rev. Biochem. (2018), 87, 479–502. https://doi.org/10.1146/annurev-biochem-062917-012550
[13] Gironda-Martínez A., Donckele E.J., Samain F., et al., DNA-Encoded Chemical Libraries: A Comprehensive Review with Successful Stories and Future Challenges, ACS Pharmacol. Transl. Sci. (2021), 4(4), 1265–1279. https://doi.org/10.1021/acsptsci.1c00118
[14] Cai B., Kim D., Akhand, S., et al., Selection of DNA-Encoded Libraries to Protein Targets within and on Living Cells, J. Am. Chem. Soc. (2019), 141(43), 17057–17061. https://doi.org/10.1021/jacs.9b08085
[15] Petersen L.K., Christensen A.B., Andersen J., et al., Screening of DNA-Encoded Small Molecule Libraries inside a Living Cell, J. Am. Chem. Soc. (2021), 143(7), 2751–2756. https://doi.org/10.1021/jacs.0c09213
[16] Huang Y., Meng L., Nie Q., et al., Selection of DNA-encoded chemical libraries against endogenous membrane proteins on live cells, Nat. Chem. (2021), 13, 77–88. https://doi.org/10.1038/s41557-020-00605-x
[17] Vipergen ApS, Understanding DEL Screening: From Target Binding to Hit Validation, Vipergen.com. Link: https://www.vipergen.com/understanding-del-screening-from-target-binding-to-hit-validation/
[18] Vipergen ApS, Molecular Glue Screening Service | DELs in Cells – Molecular Glue Direct, Vipergen.com. Link: https://www.vipergen.com/molecular-glue-direct/
[19] Molina D. M., Jafari R., Ignatushchenko M., et al., Monitoring Drug Target Engagement in Cells and Tissues Using the Cellular Thermal Shift Assay, Science (2013), 341(6141), 84–87. https://doi.org/10.1126/science.1233606
[20] Savitski M.M., Reinhard F.B.M., Franken H., et al., Tracking cancer drugs in living cells by thermal profiling of the proteome, Science (2014), 346(6205), 1255784. https://doi.org/10.1126/science.1255784
[21] Janke C., Magiera M.M., The tubulin code and its role in controlling microtubule properties and functions, Nat. Rev. Mol. Cell Biol. (2020), 21, 307–326. https://doi.org/10.1038/s41580-020-0214-3
[22] Nogales E., Wolf S.G., Khan I.A., et al., Structure of tubulin at 6.5 Å and location of the taxol-binding site, Nature (1995), 375, 424–427. https://doi.org/10.1038/375424a0
[23] Ravelli R.B.G., Gigant B., Curmi P.A., et al., Insight into tubulin regulation from a complex with colchicine and a stathmin-like domain, Nature (2004), 428, 198–202. https://doi.org/10.1038/nature02393
[24] Gigant B., Wang C., Ravelli R.B.G., et al., Structural basis for the regulation of tubulin by vinblastine, Nature (2005), 435, 519–522. https://doi.org/10.1038/nature03566
[25] Mayer T.U., Kapoor T.M., Haggarty S.J., et al., Small molecule inhibitor of mitotic spindle bipolarity identified in a phenotype-based screen, Science (1999), 286(5441), 971–974. https://doi.org/10.1126/science.286.5441.971
[26] Li Y., Zhang C., Tang D., et al., Identification of a ligand-binding site on tubulin mediating the tubulin-RB3 interaction, Proc. Natl. Acad. Sci. U.S.A. (2025), 122(11), e2424098122. https://doi.org/10.1073/pnas.2424098122
Related Services
| Service | |
|---|---|
Small molecule drug discovery for even hard-to-drug targets – identify inhibitors, binders and modulators | |
Molecular Glue Direct | |
PPI Inhibitor Direct | |
Integral membrane proteins | |
Specificity Direct – multiplexed screening of target and anti-targets | |
Express – optimized for fast turn – around-time | |
Snap – easy, fast, and affordable |


