RNA-Targeted Small Molecules and Translation-Modulating Therapeutics
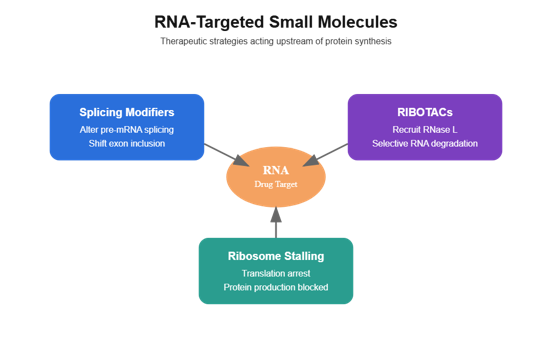
Figure: Major strategies in RNA-targeted small-molecule therapeutics. RNA-targeted pharmacology acts upstream of protein synthesis. Splicing modifiers alter exon selection in pre-mRNA to reshape transcript isoforms. RIBOTACs recruit endogenous ribonucleases such as RNase L to selectively degrade disease-causing RNA transcripts. Ribosome-stalling small molecules bind within the ribosomal exit tunnel and arrest translation of specific nascent peptide sequences, suppressing protein production. Together these approaches expand drug discovery beyond proteins to the transcriptome.
Executive Overview: Expanding the Druggable Genome Upstream of Protein
For decades, small-molecule drug discovery has focused almost exclusively on proteins — particularly enzymes and receptors with well-defined binding pockets. Yet the human genome encodes far more RNA species than druggable proteins. Messenger RNAs (mRNAs), long noncoding RNAs (lncRNAs), microRNAs, repeat-expansion transcripts, and structured untranslated regions represent a vast and largely untapped regulatory landscape.
Recent advances in structural biology, RNA chemistry, and proximity-based pharmacology have catalyzed a new frontier: RNA-targeted small molecules and translation-modulating therapeutics.
Rather than inhibiting protein function after synthesis, these strategies act upstream by:
- Modulating pre-mRNA splicing
- Degrading disease-causing RNA selectively
- Arresting translation of specific nascent polypeptides
- Exploiting RNA structure as a druggable motif
The field now encompasses:
- Splicing-modifier therapeutics
- RIBOTACs
- Ribosome-stalling small molecules (“interdictors”)
Collectively, these approaches redefine small-molecule pharmacology — from targeting static proteins to dynamically regulating gene expression itself.
Why RNA Is Emerging as a Druggable Target
The Limitations of Protein-Centric Drug Discovery
Many disease drivers are difficult to inhibit at the protein level:
- Transcription factors lacking enzymatic pockets
- Scaffolding proteins
- Gain-of-function mutations
- Proteins expressed at extremely high levels
Targeting RNA offers several advantages:
- Control before translation
- Access to noncoding regulatory molecules
- Potential for high selectivity via structural motifs
- Opportunity to eliminate toxic transcripts directly
Importantly, RNA structures are not random coils. Many RNAs fold into:
- Hairpins
- Bulges
- Internal loops
- G-quadruplexes
- Pseudoknots
These structural motifs create binding surfaces analogous to protein pockets.
Splicing-Modifying Therapeutics
Pre-mRNA splicing is a tightly regulated process that removes introns and joins exons. Errors in splicing contribute to:
- Cancer
- Neurodegeneration
- Genetic diseases
Small molecules can alter splice-site selection or spliceosome activity, shifting transcript isoforms in therapeutically beneficial ways.
Mechanistic Basis
Splicing modifiers typically interact with:
- Core spliceosome components
- RNA-protein interfaces
- Structured RNA motifs within pre-mRNA
These interactions alter exon inclusion or exclusion.
In oncology, mutations in splicing factors (e.g., SF3B1) create vulnerabilities. Cancer cells often operate near the threshold of splicing fidelity. Small perturbations can push them into transcriptomic catastrophe.
Splicing Modifiers in Cancer
Cancer cells frequently harbor:
- Spliceosome mutations
- Elevated transcriptional stress
- Increased reliance on aberrant splicing programs
Splicing modifiers can:
- Induce exon skipping in oncogenic transcripts
- Create neoantigens
- Trigger apoptosis via splicing overload
This creates a therapeutic window distinct from normal cells.
Genetic Disease Applications
Beyond oncology, splicing correction has proven transformative in inherited disorders where exon misselection drives pathology.
Small molecules offer advantages over antisense oligonucleotides:
- Oral bioavailability potential
- Broader tissue penetration
- Lower manufacturing complexity
However, transcriptome-wide effects remain a concern. Dose-dependent selectivity and careful biomarker development are critical.
RIBOTACs: Targeted RNA Degradation
RIBOTACs represent a catalytic approach to RNA elimination.
Conceptual Framework
RIBOTACs are bifunctional small molecules that:
- Bind a structured RNA target
- Recruit an endogenous ribonuclease (often RNase L)
This induced proximity triggers selective cleavage of the target RNA.
Unlike antisense oligonucleotides (ASOs) or siRNA, RIBOTACs:
- Are small molecules
- Potentially cross membranes more efficiently
- Function catalytically
- Exploit endogenous RNase machinery
Mechanistic Details
RNase L is normally activated in antiviral responses. RIBOTACs co-opt this system by tethering RNase L near a disease-relevant RNA.
Key design principles include:
- High-affinity RNA binding
- Minimal off-target RNase recruitment
- Proper geometric alignment
- Avoiding global RNase activation
The ternary complex — RNA + RIBOTAC + RNase — parallels PROTAC logic but in the RNA domain.
Comparison to Other RNA Therapeutics
| Feature | RIBOTAC | ASO | siRNA |
|---|---|---|---|
| Size | Small molecule | Oligonucleotide | Oligonucleotide |
| Delivery complexity | Moderate | High | High |
| Catalytic action | Yes | No | Yes (RISC-mediated) |
| Immune activation risk | Variable | Moderate | Moderate |
| Oral potential | Theoretical | Rare | Rare |
Target Space
Potential RNA targets include:
- Structured mRNAs
- Oncogenic lncRNAs
- Repeat expansion transcripts
- Viral RNAs
Structured motifs are particularly attractive because they create binding specificity.
Ribosome-Stalling Therapeutics (“Interdictors”)
While RIBOTACs eliminate RNA, ribosome-stalling molecules act during translation.
Mechanistic Principle
Selective ribosome-stalling compounds bind within the ribosomal exit tunnel and interact with specific nascent peptide sequences. This causes:
- Translational arrest
- Premature termination
- Reduced protein output
Importantly, stalling can be sequence-dependent, enabling selective targeting of specific proteins.
Distinguishing from Classical Translation Inhibitors
Traditional translation inhibitors (e.g., broad-spectrum antibiotics) globally suppress protein synthesis.
Ribosome-stalling therapeutics aim for:
- Target-selective arrest
- Reduced global toxicity
- Context-dependent effects
By recognizing specific amino acid motifs emerging from the ribosome, these compounds achieve precision.
Applications in Oncology
Many oncogenic drivers are difficult to degrade post-translationally due to:
- High expression
- Rapid turnover
- Structural inaccessibility
Translation arrest offers an alternative:
- Suppress production at the source
- Reduce protein accumulation
- Potentially overcome degradation resistance
Challenges
- Avoiding unintended translational stress responses
- Managing integrated stress pathway activation
- Ensuring sequence selectivity
High-resolution cryo-EM studies have been critical in elucidating binding mechanisms.
Design Principles in RNA-Targeted Small Molecules
RNA Structural Mapping
RNA structure is dynamic and context-dependent. Key tools include:
- SHAPE-MaP
- Cryo-EM
- NMR
- Chemical probing
Accurate structural models are essential for rational design.
Binding Selectivity
RNA is chemically similar across transcripts. Selectivity arises from:
- Unique structural motifs
- Bulge geometry
- Base stacking interactions
- Minor groove architecture
Medicinal chemistry must account for:
- Electrostatic interactions
- Hydrogen bonding networks
- Conformational flexibility
Avoiding Off-Target RNA Effects
Because many RNAs share structural features, off-target risks include:
- Unintended splicing shifts
- Degradation of essential transcripts
- Activation of innate immune pathways
High-throughput transcriptomic profiling is mandatory.
Induced Proximity Logic in RNA Context
Like PROTACs, RIBOTACs depend on ternary complex formation. Design parameters include:
- Orientation constraints
- Flexible vs rigid linkers
- Enzyme recruitment efficiency
- Catalytic turnover
AI-driven modeling may soon enable predictive RNA–ligand–enzyme complex design.
Therapeutic Applications
Oncology
RNA-targeted small molecules can:
- Suppress oncogenic splice variants
- Degrade oncogenic transcripts
- Arrest translation of driver proteins
- Exploit cancer-specific RNA dependencies
Tumors with splicing factor mutations are particularly vulnerable.
Repeat Expansion Disorders
Diseases caused by toxic RNA repeats (e.g., CAG, CGG expansions) are prime candidates for RIBOTAC strategies.
Selective degradation of repeat-containing transcripts could:
- Reduce toxic RNA foci
- Decrease aberrant protein translation
- Modify disease course
Viral Infections
Structured viral RNAs represent attractive targets:
- Conserved secondary structures
- Essential replication elements
- Reduced host homologs
RIBOTAC-like strategies could selectively degrade viral genomes.
Neurodegeneration
RNA-binding proteins and aberrant RNA splicing are central to:
- ALS
- Frontotemporal dementia
- Huntington’s disease
Splicing correction and RNA degradation offer novel intervention points.
Integration with Other Modalities
RNA-targeted approaches can synergize with:
- Targeted protein degradation
- Synthetic lethality strategies
- Immunotherapy
- Gene editing
For example:
- RNA suppression may sensitize tumors to degrader therapy
- Splicing modulation can create neoantigens for immune targeting
The therapeutic ecosystem is increasingly combinatorial.
Clinical and Commercial Landscape
RNA-targeted drug discovery is transitioning from exploratory chemistry to translational programs.
Key enabling factors:
- Improved RNA structural resolution
- Better predictive modeling
- Growing understanding of transcriptome dynamics
Investment interest reflects recognition that:
- RNA expands the druggable universe
- Structured RNAs provide specificity opportunities
- Small molecules offer advantages over nucleic acids in certain contexts
Companies are building pipelines around:
- Splicing modulation
- RNA degraders
- Translation regulators
Resistance Considerations
As with protein-targeted drugs, resistance may arise via:
- RNA mutation altering structure
- Alternative splicing compensation
- Upregulation of parallel pathways
- Stress response activation
Combination therapies may mitigate resistance risk.
Future Directions in RNA-Targeted Drug Discovery
The next wave of innovation may include:
- Programmable RNA degraders
- Multi-functional RNA-binding chimeras
- Tissue-specific RNA targeting strategies
- AI-guided RNA structure prediction
- Condensate-aware RNA pharmacology
Advances in computational biology will likely accelerate identification of:
- Druggable RNA motifs
- Disease-specific transcript vulnerabilities
- RNA–protein interaction hotspots
Conclusion: Rewriting the Central Dogma of Pharmacology
RNA-targeted small molecules and translation-modulating therapeutics represent a fundamental expansion of drug discovery logic.
Rather than intervening after protein synthesis, these approaches act:
- At the transcript level
- During translation
- Within splicing regulation
They convert RNA from an informational intermediary into a direct pharmacological substrate.
As the field matures, RNA-targeted drug discovery may parallel — and in some cases surpass — protein-centric approaches in impact. Structured RNAs, inducible ribonuclease recruitment, and selective ribosome stalling collectively broaden the scope of what is therapeutically addressable.
The central dogma once defined biology as DNA → RNA → protein. Modern pharmacology increasingly operates upstream — at RNA — where disease signals originate.
The era of RNA-targeted small molecules has begun.
Related Services
| Service | |
|---|---|
Small molecule drug discovery for even hard-to-drug targets – identify inhibitors, binders and modulators | |
Molecular Glue Direct | |
PPI Inhibitor Direct | |
Integral membrane proteins | |
Specificity Direct – multiplexed screening of target and anti-targets | |
Express – optimized for fast turn – around-time | |
Snap – easy, fast, and affordable |


