Unlocking the Potential of Cell-Based Assays in Modern Scientific Research
Unlocking the Potential of Cell-Based Assays in Modern Scientific Research
Introduction to Cell-Based Assays in Modern Scientific Research
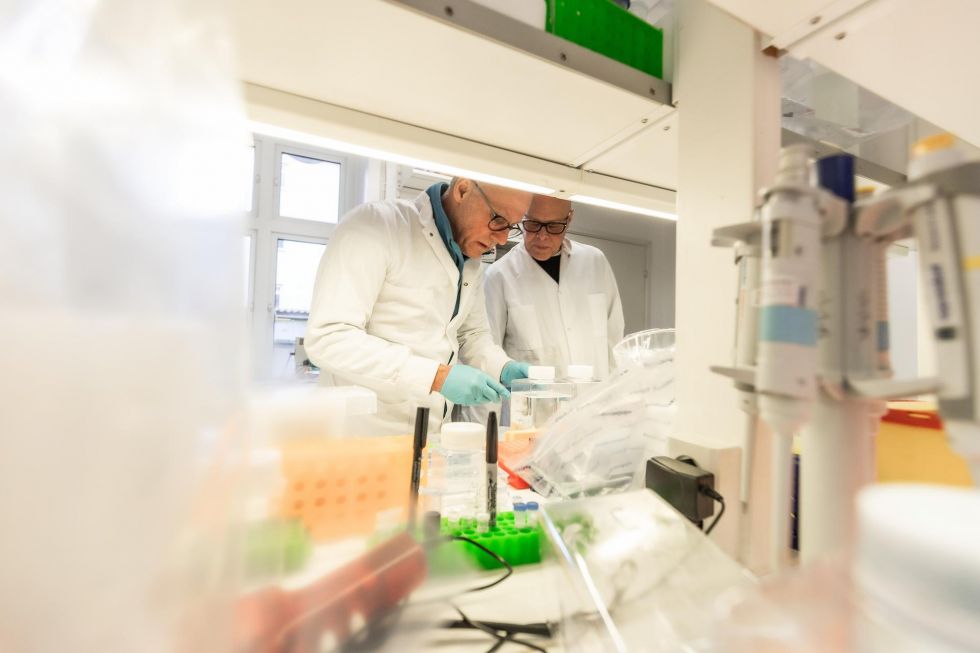
Cell-based assays play an increasingly important role in contemporary drug discovery. These assays enable researchers to evaluate the effects of chemical compounds, genetic constructs, or biological agents in living cells, under physiologically relevant environments. Cell-based assays can be applied in contexts such as early target identification, pathway elucidation, toxicity profiling and mechanistic studies. (Swinney & Anthony, 2011.)
As therapeutic targets, such as membrane-bound receptors, ion channels, and intracellular protein–protein interactions, become more complex, assay systems must adapt in parallel. This need has led to innovations in both assay design and in the integration of cell-based methods with high-throughput screening platforms.
What Are Cell-Based Assays?
Cell-based assays are experimental procedures that use live cells to assess the biochemical or physiological activities of compounds, drugs, or other biological molecules
Unlike biochemical assays, they preserve cellular context and protein interactions (Wei et al., 2021).
This enables the detection of binding events that are conditional in physiological context, providing insight into the target’s activity under near-physiological conditions. (Moffat et al.)
The defining feature of a cell-based assay is the use of intact, living cells as biosensors of molecular effects. The increasing use of Cell-based assays reflects a shift from isolated biochemical interaction measurements to dynamic observations that depend on biological context (Zhang et al., 1999).
Cell-Based vs Biochemical Assays
| Feature | Cell-Based Assays | Biochemical Assays |
|---|---|---|
| Physiological relevance | High – proteins operate in physiological relevant environment, preserve protein interaction and folding | Low – proteins studied in isolation |
| Complexity | Higher experimental complexity, more complex protocols | Relatively simple experimental setup |
| Cost | Higher due to cell culture requirements | Generally lower cost |
| Speed | Slower due to cell growth and incubation, longer development time | Faster readouts |
| Data richness | Multi-parametric cellular responses, better prediction of in vivo responses | Typically single molecular interaction |
| Reproducibility | Can vary due to biological variability | Highly reproducible |
Biochemical assays remain valuable for studying direct molecular interactions, while cell-based assays are essential for understanding how those interactions translate into biological responses. (Moffat et al., 2017).
The Role of Cell-Based Assays in Advancing Scientific Research
Cell-based assays can be categorized according to the type of detection employed (e.g., fluorescence, luminescence, impedance), the format of the assay (2D monolayers vs. 3D cultures), or the biological endpoint of interest (e.g., proliferation, signalling, cytotoxicity). (Wei et al., 2021) A wide range of assay platforms are available, each tailored to specific targets, pathways, and screening needs:
- Reporter gene assays: measure transcriptional activation using luciferase or GFP reporters.
- Cytotoxicity and Viability Assays
Assess the impact of compounds on cell survival. Common formats include ATP-based luminescence assays, resazurin reduction assays, and MTT assays.
- Cell Proliferation Assays
Measure cell division rates using markers such as BrdU incorporation or Ki-67 staining, providing insight into compound effects on growth and cell cycle progression.
- Apoptosis Assays
Detect programmed cell death through markers such as Annexin V binding, caspase activation, or DNA fragmentation.
- Calcium flux and electrophysiology assays: provide functional readouts for ion channels and GPCRs (Dunlop et al., 2008).
- Cell Migration and Invasion Assays
Assess cell motility and invasive potential, often using wound-healing assays or transwell migration systems. These assays are particularly relevant in cancer and metastasis research.
- Metabolic Activity Assays
Measure cellular metabolism using reagents such as WST-1 or resazurin to detect mitochondrial activity or metabolic conversion.
- Viability and cytotoxicity assays: use reagents such as resazurin, ATP detection, or live/dead imaging to assess cell health.
- High-content imaging (HCI/HCS): captures detailed morphological and phenotypic changes through automated microscopy (Sirenko et al., 2015).
- Oocyte-based binding assays (e.g., Vipergen’s cBTE. https://www.vipergen.com/cellular-binder-trap-enrichment/): enable DNA-tagged ligands to interact with live targets in a physiological relevant context, offering an alternative to traditional in vitro assays
Each of these methods presents specific strengths and limitations depending on the scientific question and technical setup. The table below summarizes key advantages and considerations for commonly used assay types:
Advantages and Limitations of Selected Cell-Based Assay Types
| Assay Type | Advantages | Limitations |
|---|---|---|
| Reporter gene assays | Quantitative, pathway-specific readouts; compatible with HTS formats | May not reflect post-translational regulation; artificial promoter contexts |
| Calcium flux / electrophysiology | Real-time functional data; excellent for ion channels and GPCRs | Requires specialized equipment; sensitive to variability and signal drift |
| Viability / cytotoxicity assays | Simple, scalable, and cost-effective; good for initial screening | Limited mechanistic insight; may not detect subtle cytostatic effects |
| High-content imaging (HCI/HCS) | Captures complex phenotypes; suitable for morphology, localization, etc. | Data-intensive; high analysis burden; requires image processing expertise |
| Oocyte-based binding assays (e.g. cBTE) | Preserves membrane context and protein folding; ideal for DEL screening of complex targets | Binding-only readout; requires follow-up functional assays or orthogonal data |
Selecting the appropriate assay type depends on the scientific objective, the target class, and resource availability. While viability assays may be ideal for high-throughput toxicity screens, high-content imaging and oocyte-based systems offer deeper insights into complex cellular behaviours and binding interactions, respectively.
Addressing “People Also Ask”: How Do Cell-Based Assays Work?
Cell-based assays operate by evaluating molecular interactions or biological responses in the context of intact, living cells. (Wei et al., 2021) The experimental system typically involves expressing the target protein of interest in a suitable cell type and then exposing those cells to test compounds. In some cases, the focus is on detecting compound binding under conditions that preserve physiological features such as membrane context, protein conformation, and cofactor availability.
How Cell-Based Assays Work: Basic Protocol
The experimental workflow for cell-based assays may vary depending on the biological system and assay design. However, most cell-based screening approaches follow a series of core steps that enable the evaluation of compound–target interactions in a physiologically relevant environment.
1. Cell preparation and target expression
Cells are prepared and engineered to express the target protein of interest
2. Delivery of compounds or libraries
Test compounds or compound libraries are introduced into the cellular system. In advanced screening approaches, such as DNA-encoded library (DEL) screening, compounds may be delivered together with DNA constructs that enable downstream identification.
3. Incubation and interaction
Cells are incubated under controlled conditions to allow compounds to interact with the target protein in its native cellular context. This step enables binding events that depend on proper protein folding, membrane insertion, and intracellular environment.
4. Binding stabilization and signal generation
Depending on the assay design, binding events are detected through functional readouts (e.g., fluorescence or luminescence) or through molecular enrichment strategies. In binding-based systems, conditions are applied to selectively retain compounds that interact with the target.
5. Cell lysis and sample processing
Cells are lysed to recover relevant biological material. In some workflows, additional processing steps—such as DNA ligation, amplification, or labeling—are performed to prepare samples for analysis.
6. Data collection and analysis
Signals or molecular identifiers are collected and analyzed to identify active compounds. In DEL-based approaches, next-generation sequencing is used to decode DNA barcodes associated with enriched compounds. (Gironda-Martínez et al., 2021).
7. Hit validation and follow-up
Identified hits are validated through orthogonal assays to confirm binding and assess functional activity. This step is essential to ensure biological relevance and reproducibility.
Innovative Applications and Case Studies in Cell-Based Assays
Transformative Applications in Drug Discovery and Toxicology
Modern cell-based assays have become essential tools in drug discovery by enabling the assessment of compound–target interactions in a biologically relevant context. Unlike traditional in vitro systems that rely on purified proteins or artificial membranes, cell-based assays preserve native features such as protein folding, membrane insertion, and subcellular compartmentalization. This makes them particularly valuable for screening difficult target classes like ion channels, GPCRs, and intracellular protein–protein interactions.
An important innovation in this space is the use of binding-based cell assays, which focus on compound engagement rather than downstream functional effects. For example, small molecules that act as molecular glues — stabilizing multiprotein complexes — require intact structural interfaces that are often only preserved in cellular environments. (Moffat et al) Such assays are increasingly used to validate hits from DNA-encoded library (DEL) screens and fragment libraries, helping to triage candidates based on selective and physiologically meaningful interactions.
In early toxicology, cell-based assays offer predictive insight into safety profiles. By measuring compound interactions in disease-relevant membrane environments or transport-sensitive contexts, these systems complement traditional ADMET profiling with mechanistic data on off-target binding or membrane disruption (Xu & Li, 2007).
Real-World Case Studies
- 3D cancer spheroid assays have demonstrated the ability to detect subtle cytostatic effects and morphological alterations not observable in traditional monolayer cultures, thus improving relevance for oncology models [Sirenko et al., 2015].
- High-content screening (HCS) platforms have enabled the discovery of pathway-selective modulators in complex systems such as neurodegenerative disease models, where multiparametric image analysis reveals compound-specific phenotypes beyond cell viability.
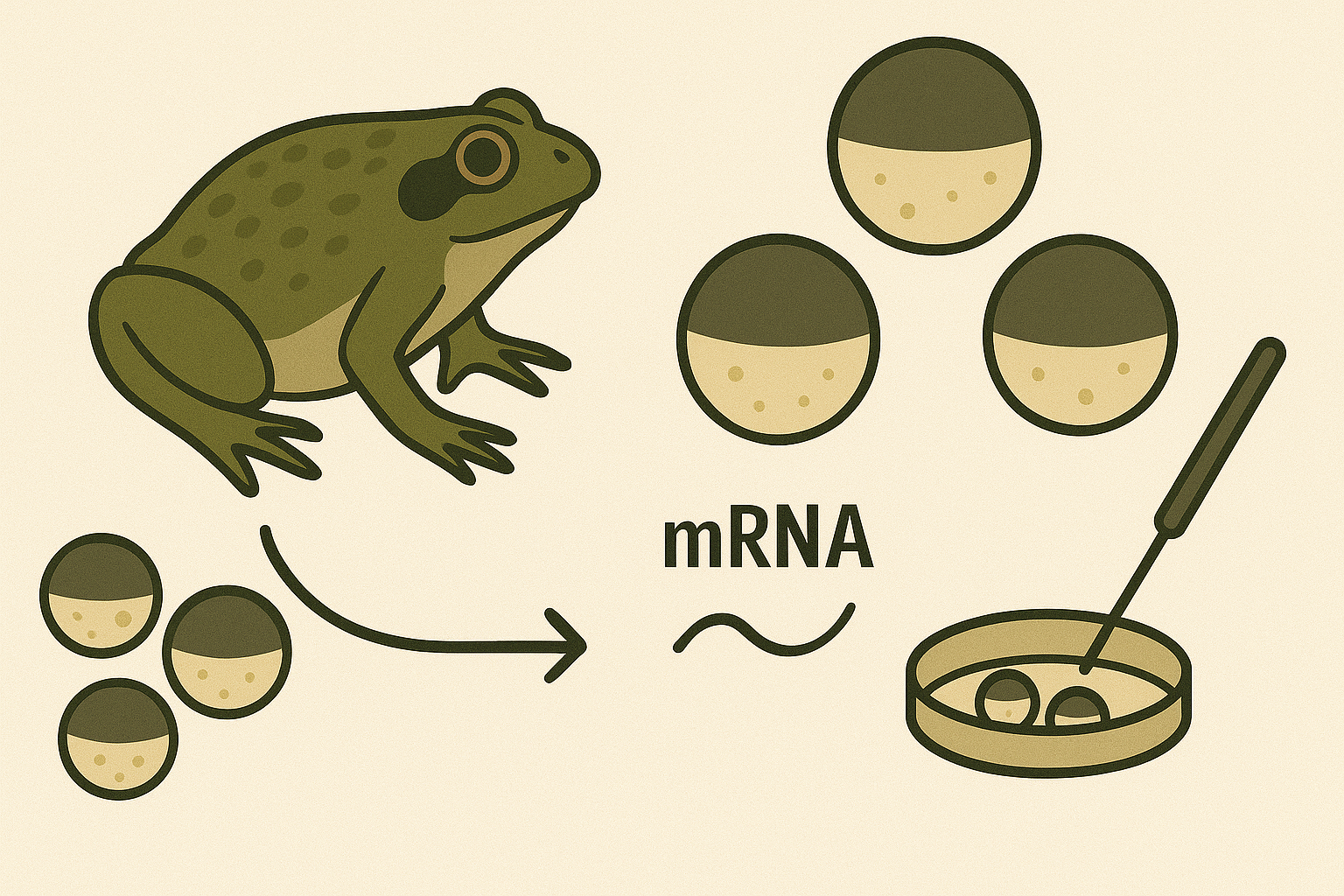
Oocyte-based binding assays, including Vipergen’s cellular Binder Trap Enrichment (cBTE) https://www.vipergen.com/cellular-binder-trap-enrichment/ screening, allow DNA-encoded libraries to be screened directly in live Xenopus laevis oocytes, a well-established system for heterologous protein expression and electrophysiological measurements (Stühmer, 1992). These cells express proteins in a native-like environment, enabling detection of binding events for structurally complex targets that are often difficult to purify or assay using classical methods.
Traditional vs. Modern Assay Techniques
While traditional assays often relied on isolated enzymes or purified receptors in artificial systems, modern cell-based platforms increasingly assess molecular interactions in a biologically relevant context. The use of intact, live cells — including oocyte-based systems — allows researchers to capture aspects of protein folding, subcellular localization, and conformational dynamics that are essential for meaningful interaction profiling.
This is particularly important in the study of challenging targets such as protein–protein interactions and molecular glues, where binding interfaces may only be preserved in native-like environments. Screening directly in living cells enables the identification of small molecules that stabilize or disrupt multiprotein complexes — an area of growing importance in next-generation drug discovery.
Emerging Trends and Innovations
Recent innovations in cell-based screening include:
- Miniaturized organoid systems for modelling tissue-specific drug responses in oncology and regenerative medicine
- Machine learning–enhanced high-content imaging, which enables phenotypic clustering and multiparametric analysis of cellular responses (Caicedo et al., 2017)
- Label-free detection technologies, such as impedance-based readouts and thermal shift assays
- Cell-based binding assays in intact membrane environments, including oocyte-based screening platforms that preserve lipid bilayers and protein folding — enabling detection of ligand interactions with GPCRs, ion channels, and protein–protein interfaces
- Assays targeting molecular glues and induced proximity, reflecting the growing interest in stabilizers of protein–protein interactions and targeted degradation mechanisms (Schapira et al., 2019)
These innovations illustrate a broader shift: from purely functional or biochemical readouts to mechanism-informed screening in biologically relevant systems.
Industry Perspectives
Robust and reproducible cell-based assays are essential for de-risking early-stage compounds. The convergence of structural biology, binding data, and cellular context has transformed how lead molecules are selected and optimized. As the field moves toward targeting previously “undruggable” proteins — such as intracellular scaffolds or dynamic complexes — the role of physiologically relevant, mechanism-specific assays becomes increasingly central to pipeline success.
For projects that must move quickly from simple binding detection to functional assays, these technologies supply chemically verified starting points already selected in a relevant cellular or solution-based context. https://www.vipergen.com/services/
Conclusion
Cell-based assays have firmly established themselves as indispensable tools in modern scientific research and drug discovery by enabling the assessment of molecular interactions in biologically relevant systems
Advances in assay formats — from 3D cultures and high-content imaging to oocyte-based binding platforms — have significantly enhanced the physiological fidelity and predictive power of these systems. This makes them especially valuable for examining challenging targets like GPCRs, ion channels, and protein–protein interactions.
As therapeutic discovery increasingly focuses on difficult and dynamic targets, the role of robust, mechanism-informed, and physiologically relevant screening platforms will only grow.
Vipergen’s contribution to cell-based screening
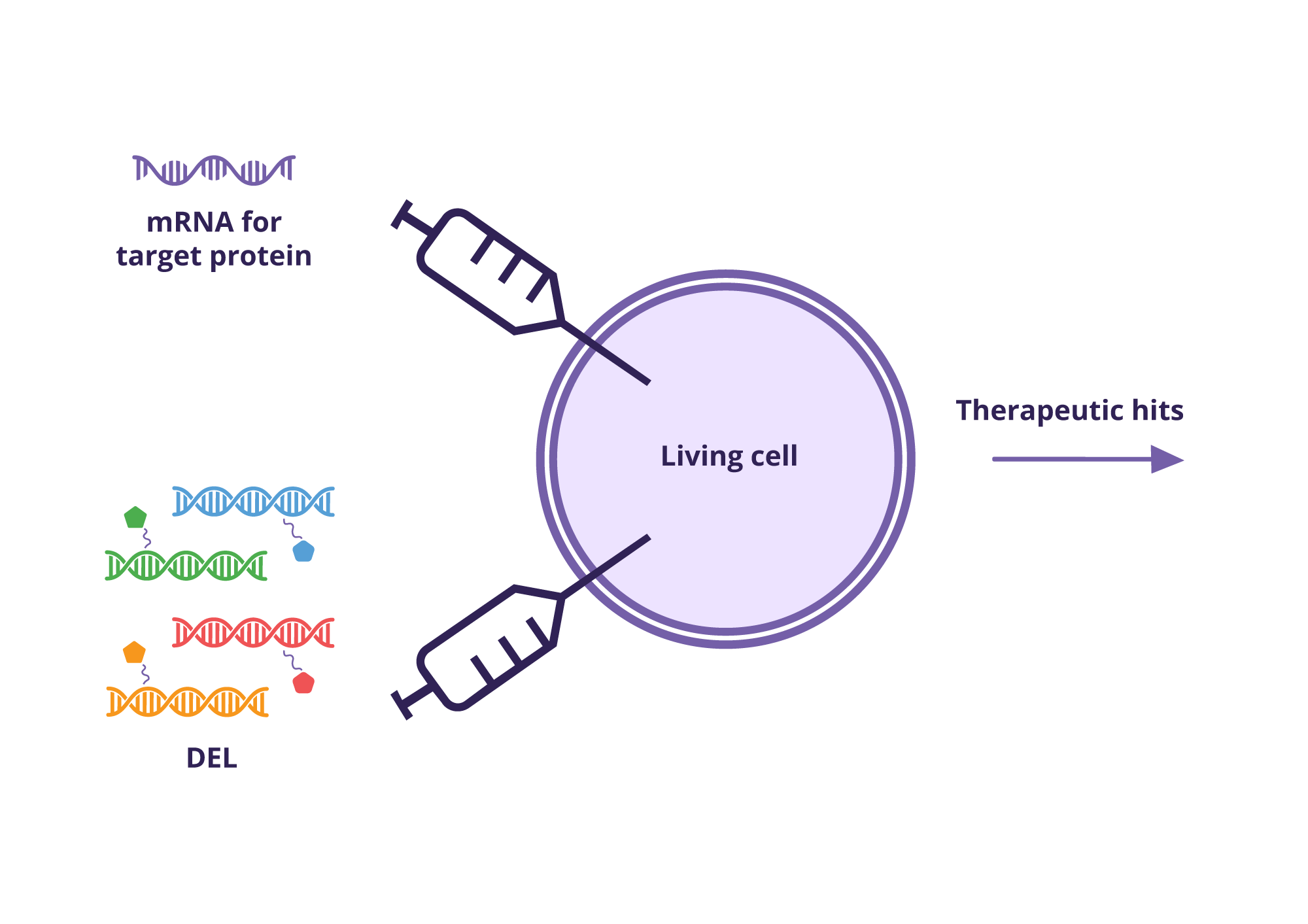
Cellular Binder Trap Enrichment (cBTE) performs the DEL screen inside living cells, so binders are identified under physiologically relevant conditions without the need for purified protein. the workflow relies on the so-called YoctoReactor libraries https://www.vipergen.com/yoctoreactor-yr-libraries/—hundreds of millions of well-defined, DNA-tagged compounds synthesised in a single tube—reducing false positives and making hit resynthesis manageable.
For projects that must move quickly from simple binding detection to functional assays, these technologies supply chemically verified starting points already selected in a relevant cellular or solution-based context. https://www.vipergen.com/services/
Frequently Asked Questions
-
What is the difference between cell-based and biochemical assays?
Cell-based assays measure how compounds and / or biological processes behave in living cells, preserving a more realistic biological context such as protein interactions, membrane environment, and cellular signaling pathways. In contrast, biochemical assays are performed on isolated molecules or proteins in vitro. Offering a mere controlled setup but less biologically relevant data.
-
What are the main types of cell-based assays?
Common types of cell-based assays include reporter gene assays, viability and cytotoxicity assays, proliferation assays, apoptosis assays, and functional assays such as calcium flux or electrophysiology. High-content imaging assays are also widely used when more detailed information about cellular behaviour or phenotype is needed.
-
How long does a cell-based assay take?
The timeline depends on assay type and biological system, but typically ranges from a few hours to several days. Assays requiring cell growth, differentiation, or longer incubation periods usually take longer than standard biochemical assays.
-
What equipment is needed for cell-based assays?
Most Cell-based assays typically require cell culture facilities, such as incubators, biosafety cabinets, and appropriate culture media. Detection systems vary depending on the assay and may include plate readers for fluorescence or luminescence, microscopes for imaging, or specialized electrophysiology equipment.
-
Why are cell-based assays important in drug discovery?
Cell-based assays play an important role in drug discovery because they show how a compound behaves in a biologically relevant environment. They provide insights into cellular responses, target engagement, and potential toxicity earlier in the process. As a result, they often provide a better indication of whether observed effects translate to in vivo systems.
References
- Swinney, D. C., & Anthony, J. (2011). How were new medicines discovered? Nat Rev Drug Discov, 10(7), 507–519. https://doi.org/10.1038/nrd3480
- Sirenko, O., Mitlo, T., Hesley, J., Luke, S., Owens, W., & Cromwell, E. F. (2015). High-content assays for characterizing the viability and morphology of 3D cancer spheroid cultures. Assay Drug Dev Technol, 13(7), 402–414. https://doi.org/10.1089/adt.2015.655
- Moffat, J. G., Vincent, F., Lee, J. A., Eder, J., & Prunotto, M. (2017). Opportunities and challenges in phenotypic drug discovery: An industry perspective. Nat Rev Drug Discov, 16(8), 531–543. https://doi.org/10.1038/nrd.2017.111
- Dunlop, J., Bowlby, M., Peri, R., Vasilyev, D., & Arias, R. (2008). High-throughput electrophysiology: An emerging paradigm for ion-channel screening and physiology. Nat Rev Drug Discov, 7(4), 358–368. https://doi.org/10.1038/nrd2552
- Stühmer, W. (1992). Electrophysiological recording from Xenopus oocytes. Methods in Enzymology, 207, 319–339. https://doi.org/10.1016/0076-6879(92)07026-
- Zhang, J.H., Chung, T.D., & Oldenburg, K.R. (1999). A Simple Statistical Parameter for Use in Evaluation and Validation of High Throughput Screening Assays. J Biomol Screen, 4(2), 67–73. https://doi.org/10.1177/108705719900400206
- Xu, J. J., Henstock, P. V., Dunn, M. C., Smith, A. R., Chabot, J. R., & de Graaf, D. (2008). Cellular imaging predictions of clinical drug-induced liver injury. Toxicological Sciences, 105(1), 97–105.
DOI: https://doi.org/10.1093/toxsci/kfn109 - Caicedo, J.C., et al. (2017). Data-analysis strategies for image-based cell profiling. Nature Methods, 14, 849–863. https://doi.org/10.1038/nmeth.4397
- Schapira, M., et al. (2019). Targeted protein degradation: expanding the toolbox. Nature Reviews Drug Discovery, 18(12), 949–963. https://doi.org/10.1038/s41573-019-0047-y
- Wei, F., Wang, S., & Gou, X. (2021). A review for cell-based screening methods in drug discovery. Biophysics Reports, 7(6), 504–516. https://doi.org/10.52601/bpr.2021.210042
- Gironda-Martínez, A., Donckele, E. J., Samain, F., Neri, D., & DNA-Encoded Chemical Libraries Network. (2021). DNA-Encoded Chemical Libraries: A Comprehensive Review with Successful Stories and Future Challenges. ACS Pharmacology & Translational Science, 4(4), 1265–1279. https://doi.org/10.1021/acsptsci.1c00118
Related Services
| Service | |
|---|---|
| Small molecule drug discovery for even hard-to-drug targets – identify inhibitors, binders and modulators | |
| Molecular Glue Direct | |
| PPI Inhibitor Direct | |
| Integral membrane proteins | |
| Specificity Direct – multiplexed screening of target and anti-targets | |
| Express – optimized for fast turn – around-time | |
| Snap – easy, fast, and affordable |